Evaluating Classification Models
Machine Learning
AI Engineering
Implementing and Evaluating Classification Models on Real-World Data

Estimated reading time: ~10 minutes
Evaluating Classification Models
Objectives
- Implement and evaluate the performance of classification models on real-world data
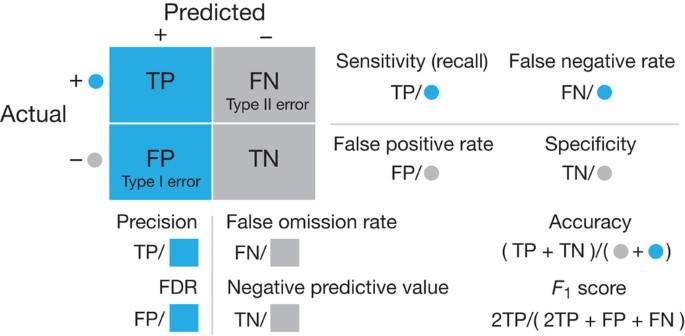
- Interpret and compare various evaluation metrics and the confusion matrix
Introduction
This article demonstrates how to use the breast cancer dataset from scikit-learn to predict whether a tumor is benign or malignant. Two classification models are created and evaluated, with Gaussian random noise added to simulate measurement errors.
Import the required libraries
import numpy as np
import pandas as pd
from sklearn.datasets import load_breast_cancer
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
from sklearn.metrics import accuracy_score, classification_report, confusion_matrix
import matplotlib.pyplot as plt
import seaborn as sns
from IPython.display import displayLoad the Breast Cancer data set
data = load_breast_cancer()
print(data.DESCR)
X, y = data.data, data.target
labels = data.target_names
feature_names = data.feature_names.. _breast_cancer_dataset:
Breast cancer wisconsin (diagnostic) dataset
--------------------------------------------
**Data Set Characteristics:**
:Number of Instances: 569
:Number of Attributes: 30 numeric, predictive attributes and the class
:Attribute Information:
- radius (mean of distances from center to points on the perimeter)
- texture (standard deviation of gray-scale values)
- perimeter
- area
- smoothness (local variation in radius lengths)
- compactness (perimeter^2 / area - 1.0)
- concavity (severity of concave portions of the contour)
- concave points (number of concave portions of the contour)
- symmetry
- fractal dimension ("coastline approximation" - 1)
The mean, standard error, and "worst" or largest (mean of the three
worst/largest values) of these features were computed for each image,
resulting in 30 features. For instance, field 0 is Mean Radius, field
10 is Radius SE, field 20 is Worst Radius.
- class:
- WDBC-Malignant
- WDBC-Benign
:Summary Statistics:
===================================== ====== ======
Min Max
===================================== ====== ======
radius (mean): 6.981 28.11
texture (mean): 9.71 39.28
perimeter (mean): 43.79 188.5
area (mean): 143.5 2501.0
smoothness (mean): 0.053 0.163
compactness (mean): 0.019 0.345
concavity (mean): 0.0 0.427
concave points (mean): 0.0 0.201
symmetry (mean): 0.106 0.304
fractal dimension (mean): 0.05 0.097
radius (standard error): 0.112 2.873
texture (standard error): 0.36 4.885
perimeter (standard error): 0.757 21.98
area (standard error): 6.802 542.2
smoothness (standard error): 0.002 0.031
compactness (standard error): 0.002 0.135
concavity (standard error): 0.0 0.396
concave points (standard error): 0.0 0.053
symmetry (standard error): 0.008 0.079
fractal dimension (standard error): 0.001 0.03
radius (worst): 7.93 36.04
texture (worst): 12.02 49.54
perimeter (worst): 50.41 251.2
area (worst): 185.2 4254.0
smoothness (worst): 0.071 0.223
compactness (worst): 0.027 1.058
concavity (worst): 0.0 1.252
concave points (worst): 0.0 0.291
symmetry (worst): 0.156 0.664
fractal dimension (worst): 0.055 0.208
===================================== ====== ======
:Missing Attribute Values: None
:Class Distribution: 212 - Malignant, 357 - Benign
:Creator: Dr. William H. Wolberg, W. Nick Street, Olvi L. Mangasarian
:Donor: Nick Street
:Date: November, 1995
This is a copy of UCI ML Breast Cancer Wisconsin (Diagnostic) datasets.
https://goo.gl/U2Uwz2
Features are computed from a digitized image of a fine needle
aspirate (FNA) of a breast mass. They describe
characteristics of the cell nuclei present in the image.
Separating plane described above was obtained using
Multisurface Method-Tree (MSM-T) [K. P. Bennett, "Decision Tree
Construction Via Linear Programming." Proceedings of the 4th
Midwest Artificial Intelligence and Cognitive Science Society,
pp. 97-101, 1992], a classification method which uses linear
programming to construct a decision tree. Relevant features
were selected using an exhaustive search in the space of 1-4
features and 1-3 separating planes.
The actual linear program used to obtain the separating plane
in the 3-dimensional space is that described in:
[K. P. Bennett and O. L. Mangasarian: "Robust Linear
Programming Discrimination of Two Linearly Inseparable Sets",
Optimization Methods and Software 1, 1992, 23-34].
This database is also available through the UW CS ftp server:
ftp ftp.cs.wisc.edu
cd math-prog/cpo-dataset/machine-learn/WDBC/
.. dropdown:: References
- W.N. Street, W.H. Wolberg and O.L. Mangasarian. Nuclear feature extraction
for breast tumor diagnosis. IS&T/SPIE 1993 International Symposium on
Electronic Imaging: Science and Technology, volume 1905, pages 861-870,
San Jose, CA, 1993.
- O.L. Mangasarian, W.N. Street and W.H. Wolberg. Breast cancer diagnosis and
prognosis via linear programming. Operations Research, 43(4), pages 570-577,
July-August 1995.
- W.H. Wolberg, W.N. Street, and O.L. Mangasarian. Machine learning techniques
to diagnose breast cancer from fine-needle aspirates. Cancer Letters 77 (1994)
163-171.
Standardize the data and add noise
scaler = StandardScaler()
X_scaled = scaler.fit_transform(X)
np.random.seed(42)
noise_factor = 0.5
X_noisy = X_scaled + noise_factor * np.random.normal(loc=0.0, scale=1.0, size=X.shape)
df = pd.DataFrame(X_scaled, columns=feature_names)
df_noisy = pd.DataFrame(X_noisy, columns=feature_names)
print("riginal Data (First 5 rows):")
display(df.head())
print("nNoisy Data (First 5 rows):")
display(df_noisy.head())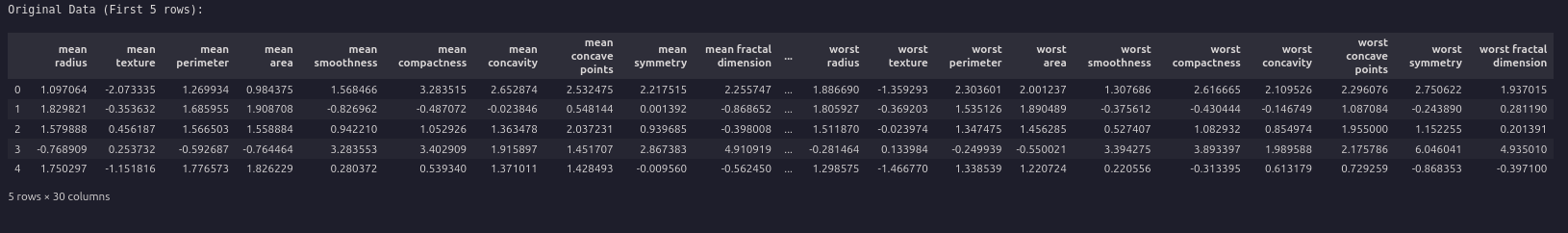
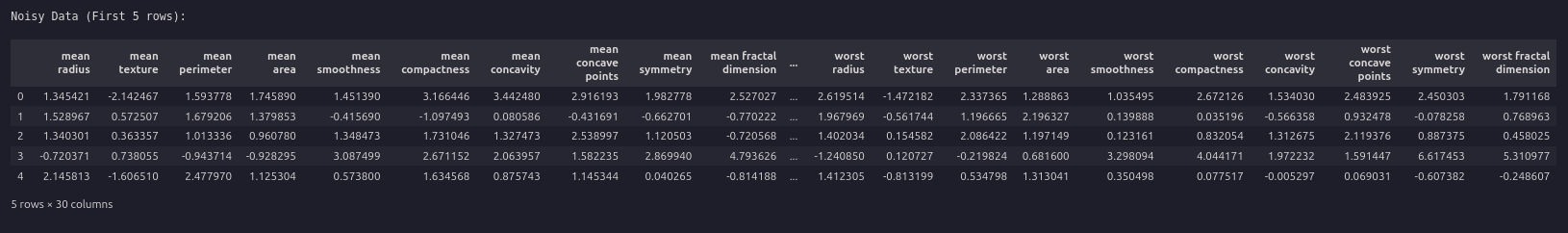
Split the data and fit KNN and SVM models
X_train, X_test, y_train, y_test = train_test_split(X_noisy, y, test_size=0.3, random_state=42)
knn = KNeighborsClassifier(n_neighbors=5)
svm = SVC(kernel='linear', C=1, random_state=42)
knn.fit(X_train, y_train)
svm.fit(X_train, y_train)Visualize the noise content
We can get a good idea of how much noise there is in the features by comparing values in the previous tables. We can also visualize the differences in several ways. Let’s begin by plotting the histograms of one of the features with and without noise for comparison.
plt.figure(figsize=(12, 6))
# Original Feature Distribution (Noise-Free)
plt.subplot(1, 2, 1)
plt.hist(df[feature_names[5]], bins=20, alpha=0.7, color='blue', label='Original')
plt.title('Original Feature Distribution')
plt.xlabel(feature_names[5])
plt.ylabel('Frequency')
# Noisy Feature Distribution
plt.subplot(1, 2, 2)
plt.hist(df_noisy[feature_names[5]], bins=20, alpha=0.7, color='red', label='Noisy')
plt.title('Noisy Feature Distribution')
plt.xlabel(feature_names[5])
plt.ylabel('Frequency')
plt.tight_layout() # Ensures proper spacing between subplots
plt.show()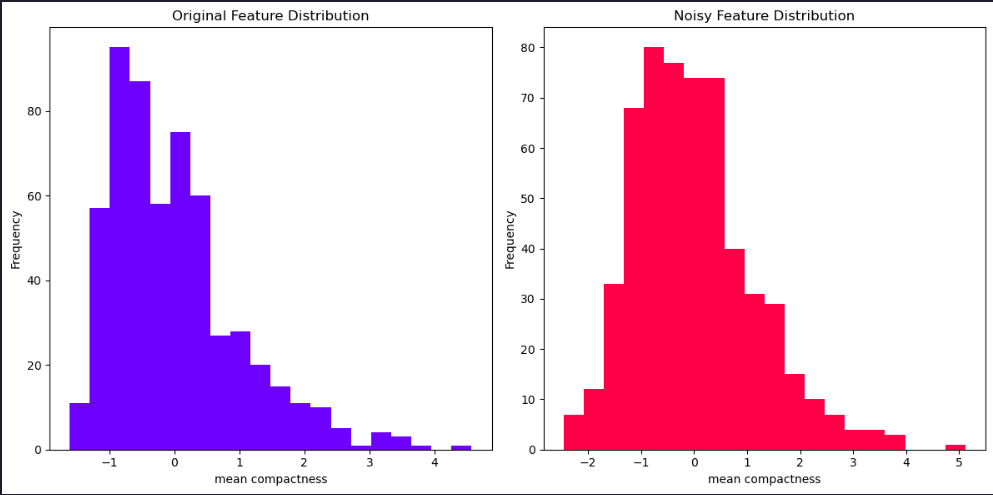
plt.figure(figsize=(12, 6))
plt.plot(df[feature_names[5]], label='Original',lw=3)
plt.plot(df_noisy[feature_names[5]], '--',label='Noisy',)
plt.title('Scaled feature comparison with and without noise')
plt.xlabel(feature_names[5])
plt.legend()
plt.tight_layout()
plt.show()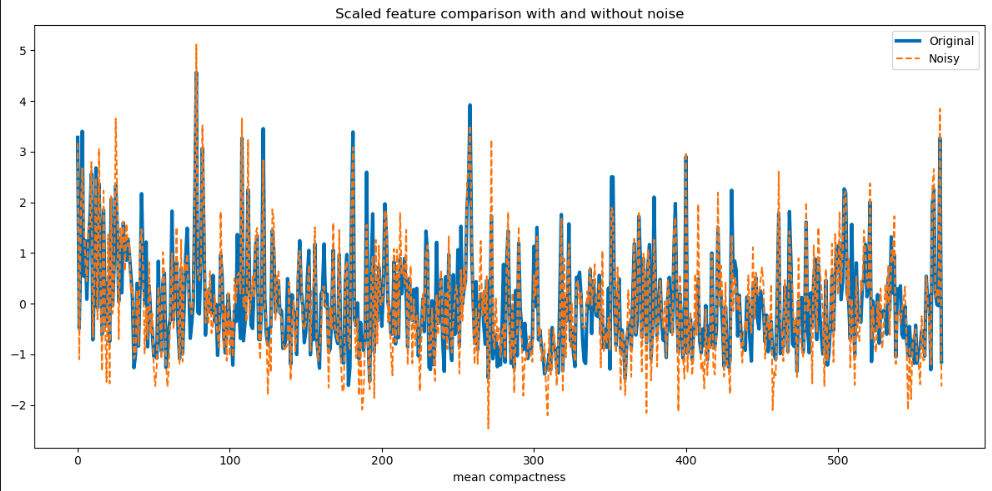
plt.figure(figsize=(12, 6))
plt.scatter(df[feature_names[5]], df_noisy[feature_names[5]],lw=5)
plt.title('Scaled feature comparison with and without noise')
plt.xlabel('Original Feature')
plt.ylabel('Noisy Feature')
plt.tight_layout()
plt.show()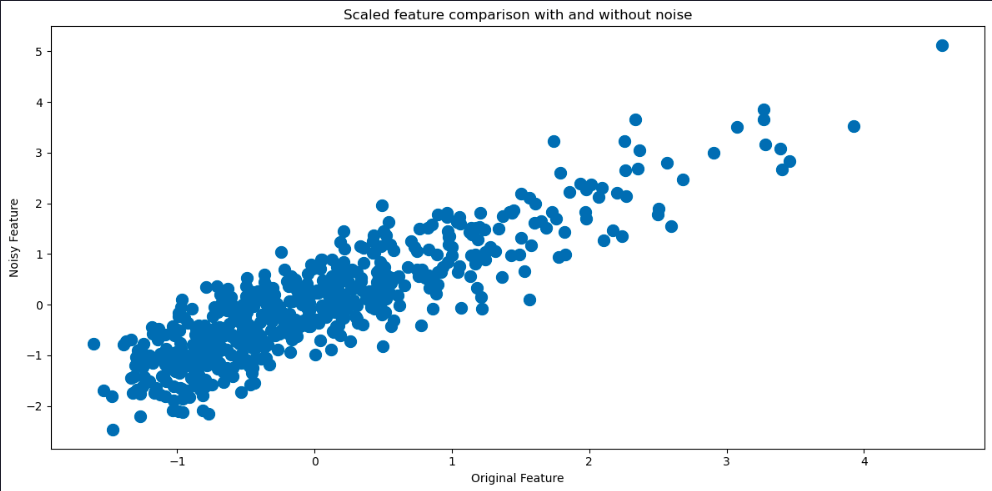
Evaluate the models
y_pred_knn = knn.predict(X_test)
y_pred_svm = svm.predict(X_test)
print(f"KNN Testing Accuracy: {accuracy_score(y_test, y_pred_knn):.3f}")
print(f"SVM Testing Accuracy: {accuracy_score(y_test, y_pred_svm):.3f}")
print("\nKNN Testing Data Classification Report:")
print(classification_report(y_test, y_pred_knn))
print("\nSVM Testing Data Classification Report:")
print(classification_report(y_test, y_pred_svm))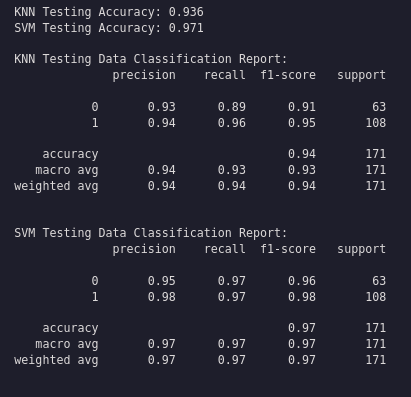
Plot the confusion matrices
conf_matrix_knn = confusion_matrix(y_test, y_pred_knn)
conf_matrix_svm = confusion_matrix(y_test, y_pred_svm)
fig, axes = plt.subplots(1, 2, figsize=(12, 5))
sns.heatmap(conf_matrix_knn, annot=True, cmap='Blues', fmt='d', ax=axes[0], xticklabels=labels, yticklabels=labels)
axes[0].set_title('KNN Testing Confusion Matrix')
axes[0].set_xlabel('Predicted')
axes[0].set_ylabel('Actual')
sns.heatmap(conf_matrix_svm, annot=True, cmap='Blues', fmt='d', ax=axes[1], xticklabels=labels, yticklabels=labels)
axes[1].set_title('SVM Testing Confusion Matrix')
axes[1].set_xlabel('Predicted')
axes[1].set_ylabel('Actual')
plt.tight_layout()
plt.show()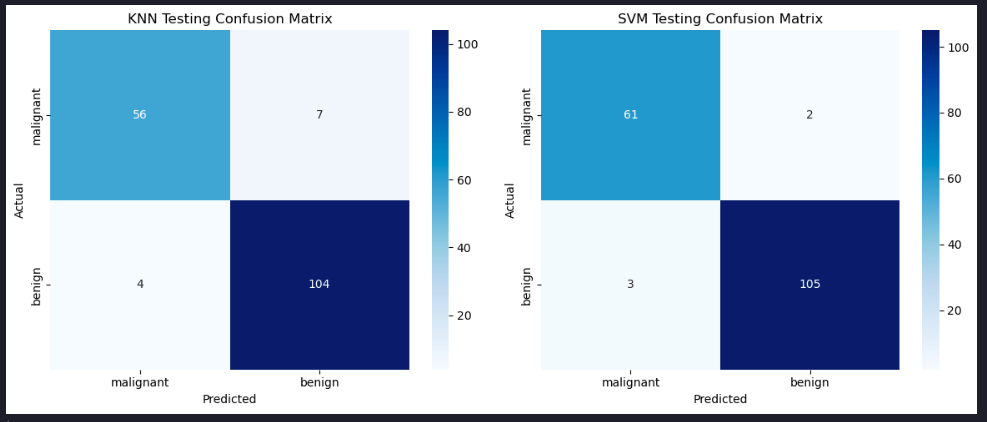
Summary
This article demonstrated how to implement and assess the performance of classification models using real-world data, exploring different evaluation metrics and the confusion matrix.